3.6 Are all Lawsonia intracellularis isolates the same?
PE has been reported in a broad host range, including pigs, non-human primates, hamsters, rabbits, rats, guinea pigs, foals, sheep, white-tailed deer, ferrets, arctic foxes, dogs and certain birds (Cooper and Gebhart 1998). L. intracellularis infection has been established experimentally in mice that are deficient in interferon-gamma receptors and has been reported to occur spontaneously in conventional mice. There are no reports of PE in human beings, but recent reports of the disease in non-human primates suggests that, as diagnostic methods improve, the disease may be detected in humans.
The source of L. intracellularis in affected animal species has not been determined. It may be endemic in certain species or it may be present in the environment. Species to species transmission of L. intracellularis has been documented. Pig isolates can infect hamsters, mice and horses and a horse isolate can infect hamsters (Cooper and Gebhart 1998; AlGhamdi 2003).
Feces from infected pigs may provide the source of new infections for susceptible animals (McOrist and Gebhart 1999) and pig-to-pig contact has been shown to be an important route of transmission. Other possible mechanisms of transmission of L. intracellularis include mechanical vectors, such as rubber boots, and biological vectors, such as mice, small birds and insects.
Characterization of Lawsonia isolate differences is hindered by difficulty in its primary cultivation and identification. Phylogenetic analysis and comparison of the 16S rDNA sequences of the intracellular organism obtained from PE in various animal species were performed. Results indicate that the intracellular agents from these species have high sequence similarity, indicating that they are all in the genus Lawsonia and that they may also be the same species, L. intracellu-laris. Due to the close similarity of the 16S rDNA of Lawsonia isolates obtained from these diverse animal species, 16S rDNA-based molecular epidemiology techniques is not able to characterize differences in L. intracellularis isolates.
Analysis of the 16S-23S rDNA intergenic spacer region of bacteria may discriminate between isolates which have identical 16S rDNA sequences. Primers that specifically amplify the 16S to 23S intergenic spacer region of L. intracellularis and related organisms (Bilophila wadsworthia) were designed. The sequence obtained from these primers with the type strain of L. intracellu-laris was identical in both length and sequence to those obtained from pure culture L. intra-cellularisintracellularis isolates from pigs with both forms of PE and obtained from the U.S., the U.K., and Denmark. The fact that identical PCR products were obtained from all Lawsonia intracellularis isolates suggests that all isolates tested are genetically similar. The sequence of Lawsonia’s closest relative, Bilophila wadsworthia, differed over a similar region by more than 40% of the bases.
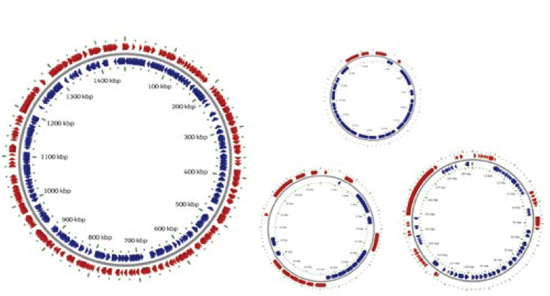
Figure 3.6 a
Lawsonia intracellularis Genome + 3 plasmids
1,719,350 bp;
1,346 ORFs
The complete
genome of Lawsonia intracellularis has recently been decoded.
The outer-membrane protein (OMP) profiles of six different L. intracellularis isolates and Bilophila wadsworthia were examined by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). The sarcosinate-insoluble membrane preparations of all six Lawsonia intracellularis isolates had the same pattern of six major bands, which were 78, 48, 37, 28, 23, and 17.6 Kb. B. wadsworthia had four major bands that were at 47, 45, 37, and 17.6 Kb. All six L. intra-cellularis isolates had the same OMP profile and two of the four B. wadsworthia OMP bands were similar in size to L. intracellularis OMP bands. Based on the OMP profile using SDS-PAGE, all six Lawsonia intracellularis isolates were phenotypically similar.
Short sequence repeats or variable number tandem repeats (VNTR) have been found in non-coding parts of the genome of several bacterial species and have been used for subspecific differentiation of isolates. VNTRs were found in the LI genome and their utility as markers for differentiating isolates of Lawsonia intracellularis was demonstrated. Analysis of VNTR profiles can be a useful epidemiological tool for distinguishing between isolates of L. intracellularis around the world. The assay proved to be robust and gave identical results on repeat analysis.
© Boehringer Ingelheim Animal Health GmbH, 2006
All rights reserved. No part of this Technical Manual 3.0 may be reproduced or transmitted in any form or by any means, electronic or photocopy, without permission in writing from Boehringer Ingelheim Animal Health GmbH.




